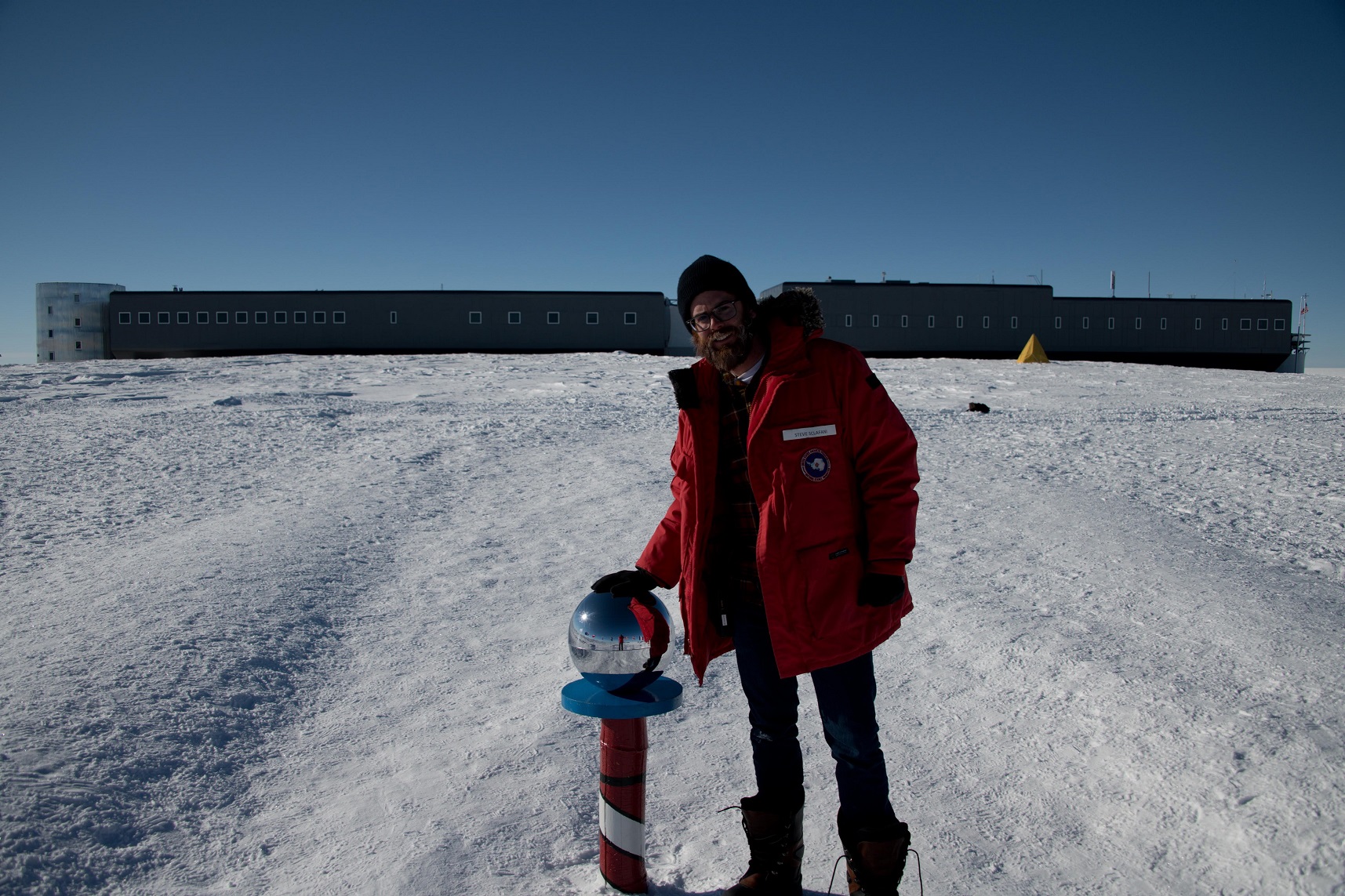
Steve Sclafani at the South Pole.
Steve Sclafani at the South Pole. Breakthrough Prize Awarded to CERN Experiments
- Details
- Category: Department News
- Published: Monday, April 07 2025 01:40
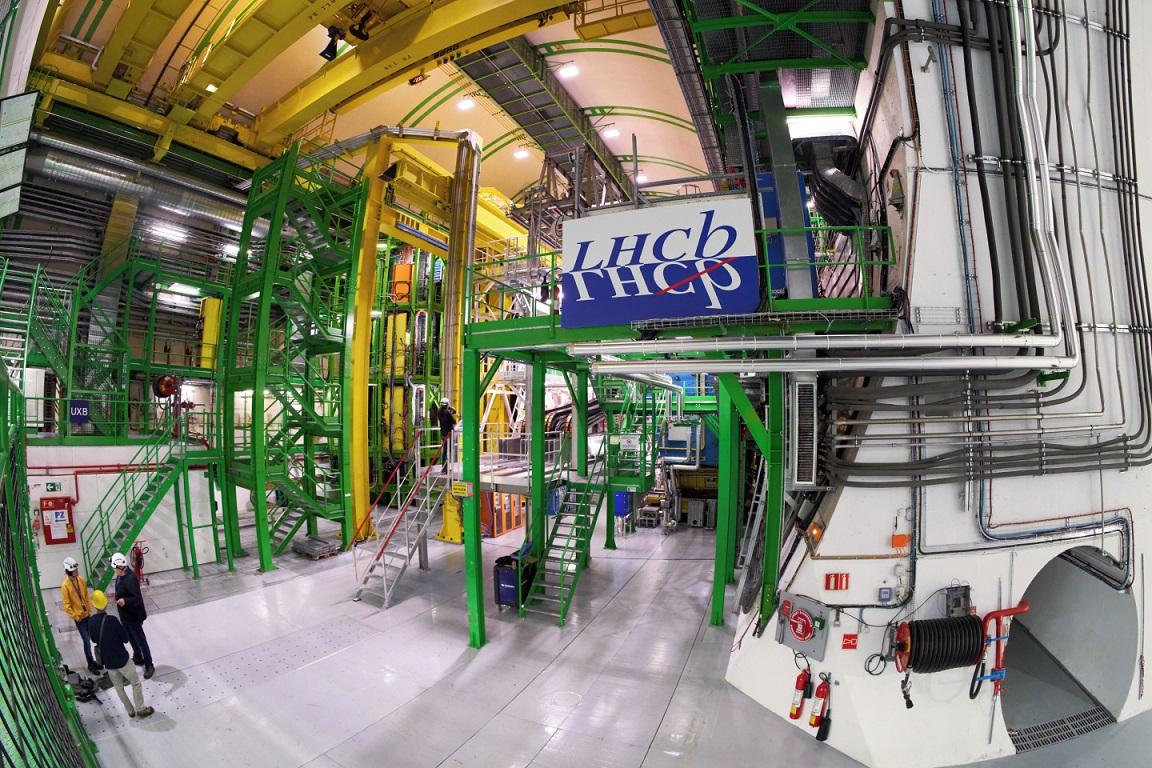
On April 5, 2025, the CMS, LHCb, ALICE and ATLAS collaborations at the Large Hadron Collider at CERN were honored with the Breakthrough Prize in Fundamental Physics by the Breakthrough Prize Foundation. The prize is awarded to the four collaborations, which unite thousands of researchers from more than 70 countries, and concerns the papers authored based on LHC Run-2 data up to July 2024. It was received by the spokespersons who led the collaborations during that time.
The prize was awarded to the collaborations for their “detailed measurements of Higgs boson properties confirming the symmetry-breaking mechanism of mass generation, the discovery of new strongly interacting particles, the study of rare processes and matter-antimatter asymmetry, and the exploration of nature at the shortest distances and most extreme conditions at CERN’s Large Hadron Collider”.
“I am extremely proud to see the extraordinary accomplishments of the LHC collaborations honoured with this prestigious Prize,” said Fabiola Gianotti, Director-General of CERN. “It is a beautiful recognition of the collective efforts, dedication, competence and hard work of thousands of people from all over the world who contribute daily to pushing the boundaries of human knowledge.”
Following consultation with the experiments’ management teams, the Breakthrough Prize Foundation will donate the $3 million Prize to the CERN & Society Foundation. The Prize money will be used to offer grants for doctoral students from the collaborations’ member institutes to spend research time at CERN, giving them experience in working at the forefront of science and new expertise to bring back to their home countries and regions.
Many current and past UMD physicists have contributed to the CMS and LHCb experiments. During run 2, Alberto Belloni was project leader for the CMS HCAL, and Sarah Eno was USCMS Collaboration board deputy chair.
ATLAS and CMS are general-purpose experiments, which pursue the full programme of exploration offered by the LHC’s high-energy and high-intensity proton and ion beams. They jointly announced the discovery of the Higgs boson in 2012 and continue to investigate its properties.
"This prize recognises the collective vision and monumental effort of thousands of ATLAS collaborators worldwide", says ATLAS spokesperson Stephane Willocq. "Their talent and dedication, and the support of our public funding agencies, enabled the scientific breakthroughs that are being celebrated today. These results have transformed our understanding of the Universe at the most fundamental level.”
"CMS is deeply honoured to receive this prestigious prize,” said CMS spokesperson Gautier Hamel de Monchenault. “Through continuous innovation in exploiting the data from the Large Hadron Collider over the past fifteen years, the CMS collaboration is conducting a thorough characterisation of the Higgs boson, exploring the electroweak scale and beyond and probing the hot, dense state of nuclear matter that prevailed in the early Universe.”
ALICE studies quark-gluon plasma, a state of extremely hot and dense matter that existed in the first microseconds after the Big Bang, while LHCb explores minute differences between matter and antimatter, violation of fundamental symmetries and the complex spectra of composite particles (“hadrons”) made of heavy and light quarks, among other things.
“The ALICE collaboration is honoured to receive the Breakthrough Prize for the investigation of the properties of the hottest and densest matter available in a laboratory, quark-gluon plasma”, says ALICE spokesperson Marco Van Leeuwen. “The new grants funded through this prize will contribute to training the next generation of ALICE scientists.”
"The award of the 2025 Breakthrough Prize is a great honour for the LHCb collaboration. It underlines the importance of the many measurements made by the LHCb experiment in flavour physics and spectroscopy through the exploration of subtle differences between matter and antimatter and the discovery of several new heavy quark hadrons”, says LHCb spokesperson Vincenzo Vagnoni.
By performing these extraordinarily precise and delicate tests, the LHC experiments have pushed the boundaries of knowledge of fundamental physics to unprecedented limits. They will continue to do so with the upcoming upgrade of the Large Hadron Collider, the High-Luminosity LHC, which aims to ramp up the performance of the LHC, starting in 2030, in order to increase the potential for discoveries.
Original story: https://home.cern/news/press-release/knowledge-sharing/lhc-experiment-collaborations-cern-receive-breakthrough-prize